Professor Bertie Göttgens

Professor Bertie Göttgens
Cellular decision making in normal and leukaemic blood stem cells
Email: bg200@cam.ac.uk | Departmental Affiliation: Haematology
The Göttgens Lab is jointly led by Professor Bertie Göttgens(Director of the Cambridge Stem Cell Institute) and Research Professor Nicola Wilson. Together, they guide the lab’s efforts to understand the gene regulatory networks that control blood stem cell function and their disruption in disease.
Research
Plain English:Blood stem cells maintain the lifelong production of all blood cell types. This depends on finely tuned control systems which can fail as we age or go wrong and lead to disease (leukaemia). The Göttgens Group uses experimental and computational approaches to discover how these control systems work and how their disruption leads to disease.
Research Focus: Combining experimental and computational approaches, the Göttgens group has mapped the gene regulatory networks that control blood stem cells and revealed how their disruption contributes to leukaemia. More recently, the group has harnessed single-cell and time-series genomics to build dynamic models of blood production, revealing how stem cells adapt during development, ageing and disease.
Current research focuses on:
(i) early blood development from pluripotent cells
(ii) mechanisms of cellular decision making in blood stem and progenitor cells
(iii) functional consequences of leukaemogenic mutations
(iv) computational modelling of normal and perturbed haematopoiesis

Göttgens Group photo
Funding
MRC, Wellcome, Blood Cancer UK, Cancer Research UK, NIH, Aging Biology Foundation
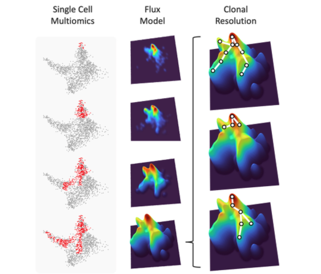
From single cell time-series to clonally resolved cellular flux models: The three columns illustrate the progression from time-resolved single cell haematopoietic landscapes to cellular flux models, ultimately drilling down to capturing the collective behaviour of individual stem cell clones.
Key Publications
- Theeuwes B, Harland LTG, Bisia AM, Costello I, Ton MN, Lohoff T., Clark SJ, Argelaguet R., Wilson NK, Wolf Reik W., Bikoff EK, Robertson EJ, Göttgens B. (2025) Eomes directs the formation of spatially and functionally diverse extraembryonic hematovascular tissuesDevelopmental Cell DOI: 10.1016/j.devcel.2025.06.001
- Dellorusso PV, Proven MA, Calero-Nieto FJ, Wang X, Mitchell CA, Hartmann F, Amouzgar M, Favaro P, DeVilbiss A, Swann JW, Ho TT, Zhao Z, Bendall SC, Morrison S, Göttgens B, Passegué E. (2024) Autophagy counters inflammation-driven glycolytic impairment in aging hematopoietic stem cells. Cell Stem Cell. 2024 Jul 5;31(7):1020-1037.e9
- Isobe T, Kucinski I, Barile M, Wang X, Hannah R, Bastos HP, Chabra S, Vijayabaskar MS, Sturgess KHM, Williams MJ, Giotopoulos G, Marando L, Li J, Rak J, Gozdecka M, Prins D, Shepherd MS, Watcham S, Green AR, Kent DG, Vassiliou GS, Huntly BJP, Wilson NK, Göttgens B. (2023) Preleukemic single-cell landscapes reveal mutation-specific mechanisms and gene programs predictive of AML patient outcomesCell Genomics 2023 Oct 27;3(12):100426
- Kucinski, I., Campos, J., Barile, M., Severi, F., Bohin, N., Moreira, P.N., Allen, L., Lawson, H., Haltalli, M.L.R., Kinston, S.J., O’Carroll, D., Kranc, K.R., Göttgens, B. (2024) A time and single-cell resolved model of murine bone marrow hematopoiesis. Cell Stem Cell. 31(2): 244-259
- Jardine L, Webb S, Goh I, Quiroga Londoño M, Reynolds G, Mather M, Olabi B, Stephenson E, Botting RA, Horsfall D, Engelbert J, Maunder D, Mende N, Murnane C, Dann E, McGrath J, King H, Kucinski I, Queen R, Carey CD, Shrubsole C, Poyner E, Acres M, Jones C, Ness T, Coulthard R, Elliott N, O'Byrne S, Haltalli MLR, Lawrence JE, Lisgo S, Balogh P, Meyer KB, Prigmore E, Ambridge K, Jain MS, Efremova M, Pickard K, Creasey T, Bacardit J, Henderson D, Coxhead J, Filby A, Hussain R, Dixon D, McDonald D, Popescu DM, Kowalczyk MS, Li B, Ashenberg O, Tabaka M, Dionne D, Tickle TL, Slyper M, Rozenblatt-Rosen O, Regev A, Behjati S, Laurenti E, Wilson NK, Roy A, Göttgens B*, Roberts I*, Teichmann SA*, Haniffa M*. Blood and immune development in human fetal bone marrow and Down syndrome. Nature. 2021 Oct;598(7880):327-331. PMCID: PMC7612688
- Stephenson E, Reynolds G, Botting R.A., Calero-Nieto F.J., Morgan M.D., Tuong Z.K.,. Bach K, Sungnak W, Worlock KB, Yoshida M, Kumasaka N, Kania K, Engelbert J, Olabi B, Spegarova JS, Wilson NK … Smith K.G.C., Teichmann S.A., Clatworthy M.R., Marioni J.C., Göttgens B*, Haniffa M. Single-cell multi-omics analysis of the immune response in COVID-19. Nat Med. 2021 Apr 20. doi: 10.1038/s41591-021-01329-2. PMCID: PMC8121667
- Pijuan-Sala B., Wilson N.K., Xia J., Hou X., Hannah R.L., Kinston S., Calero-Nieto F.J., Poirion O., Preissl S., Liu F., Göttgens B. (2020). Single-Cell Chromatin Accessibility Maps Reveal Regulatory Programs Driving Early Mouse Organogenesis Nature Cell Biol 22: 487-497 PMCID: PMC7145456
- Kucinski I., Wilson N.K., Hannah R., Kinston S.J., Cauchy P., Lenaerts A., Grosschedl R., Göttgens B. (2020). Cooperation of Lineage-associated Transcription Factors Governs Haematopoietic Progenitor States EMBO Journal 39: e104983 PMCID: PMC7737608
- Pijuan-Sala B, Griffiths JA, Guibentif C, Hiscock TW, Jawaid W, Calero-Nieto FJ, Mulas C, Ibarra-Soria X, Tyser RCV, Ho DLL, Reik W, Srinivas S, Simons BD, Nichols J, Marioni JC, Göttgens B, (2019) A single-cell molecular map of mouse gastrulation and early organogenesis. Nature 566 (7745) 490-495 PMCID: PMC6522369
- Lescroart F, Wang X, Lin X, Swedlund B, Gargouri S, Sànchez-Dànes A, Moignard V, Dubois C, Paulissen C, Kinston S, Göttgens B*, Blanpain C. Defining the earliest step of cardiovascular lineage segregation by single-cell RNA-seq. Science. 2018 Mar 9;359(6380):1177-1181. *
- Ibarra-Soria X, Jawaid W, Pijuan-Sala B, Ladopoulos V, Scialdone A, Jörg DJ, Tyser RCV, Calero-Nieto FJ, Mulas C, Nichols J, Vallier L, Srinivas S, Simons BD, Göttgens B*, Marioni JC. Defining murine organogenesis at single-cell resolution reveals a role for the leukotriene pathway in regulating blood progenitor formation. Nat Cell Biol. 2018 Feb;20(2):127-134. *
- Scialdone A., Tanaka Y., Jawaid W., Moignard V., Wilson N.K., Macaulay I.C., Marioni J.C., Göttgens B. (2016). Resolving Early Mesoderm Diversification through Single Cell Expression Profiling Nature 535 (7611): 289-29
- Wilson NK, Kent DK, Buettner F, Shehata M, Macaulay IC, Calero-Nieto FJ, Sánchez Castillo M, Oedekoven CA, Diamanti E, Schulte R, Ponting CP, Voet T, Caldas C, Stingl J, Green AR, Theis FJ, Göttgens B. (2015). Combined single cell functional and gene expression analysis resolves heterogeneity within stem cell populationsCell Stem Cell 16(6): 712-724. PMCID:PMC4460190
* joint senior author
