Dr Brian Hendrich
Transcriptional control of stem cell fate
Email: bdh24@cam.ac.uk
Laboratory: Cambridge Stem Cell Institute, Jeffrey Cheah Biomedical Centre
Departmental Affiliation: Biochemistry
Biography
Brian Hendrich received his PhD from Stanford University in 1995 working on X chromosome inactivation with Huntington Willard. In 1995 he joined the lab of Adrian Bird at the University of Edinburgh and participated in the discovery and characterisation of a family of methyl-CpG binding proteins in mammals. In 2001 he started his own laboratory at the University of Edinburgh. In 2008 he moved to the Wellcome Trust Centre for Stem Cell Research in Cambridge.
He is currently a Principal Research Associate and Director of the PhD Programme in Stem Cell Biology and Medicine for the Wellcome - MRC Cambridge Stem Cell Institute.
Funding
Wellcome, Medical Research Council, Wolfson College, Astra Zeneca
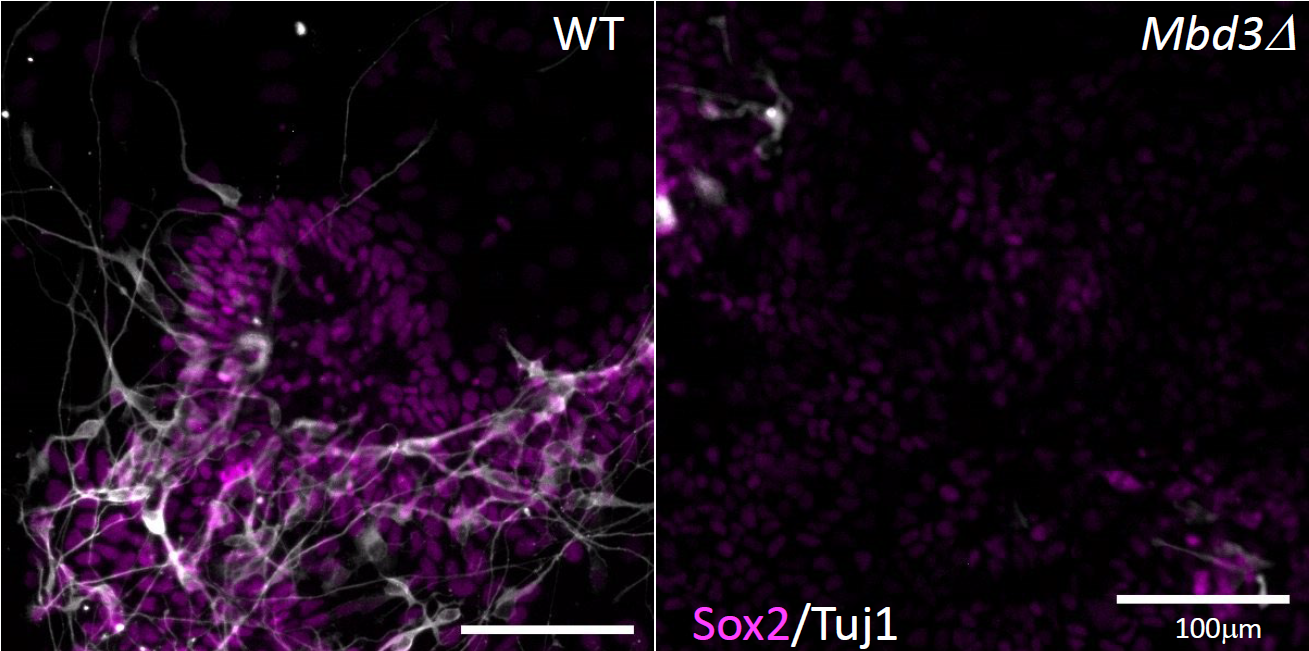
EpiStem Cells form neurons (white) when differentiated in culture (left). In the absence of Mbd3/NuRD (right) cells are unable to undergo neural differentiation.
Research
Embryonic stem (ES) cells hold enormous promise for personalised medicine and drug discovery since they can be maintained indefinitely and are pluripotent. While pluripotency makes ES cells potentially very useful, it also presents a problem: how do you get them to make the cell type you want, and not one you don’t? Differentiation of pluripotent cells is exquisitely organised during normal embryogenesis, but very hard to control in culture.
Since all cells in an organism are genetically identical, the observable differences in their functions and behaviours come down to which genes they express and which genes they repress. In order to understand how to direct cellular identity, our lab seeks to understand how cells execute the changes in gene expression control needed to successfully and appropriately respond to differentiation signals.
By understanding how ES cells make different developmental decisions, the lab hopes to bring the medical promise of stem cells closer to realisation.
Hendrich Group photo 2023
Plain English
Embryonic stem cells can either make more copies of themselves, or differentiate to form any cell type in the body. This means that they have the potential to form any tissue, that is, they are pluripotent. How pluripotent cells make the decision to differentiate, and which cell type to differentiate into, is defined by which genes the cell turns on, and which it turns off. Genes are encoded in the cell’s DNA, which gets packaged up in the cell with proteins into a structure called chromatin. My group studies the function of a group of proteins which can change the structure of chromatin, thereby turning genes up, down, or off, in pluripotent cells. Our key questions are: How do pluripotent cells control their gene expression in order to make developmental decisions? How does the function of chromatin modifying proteins precisely control gene expression patterns? We are addressing these questions by studying both embryonic stem cells, but also pluripotent cells that exist very early in mammalian development. We aim to better understand how cells make decisions during normal development, but also to understand how these processes occasionally go wrong and result in human diseases such as cancer.
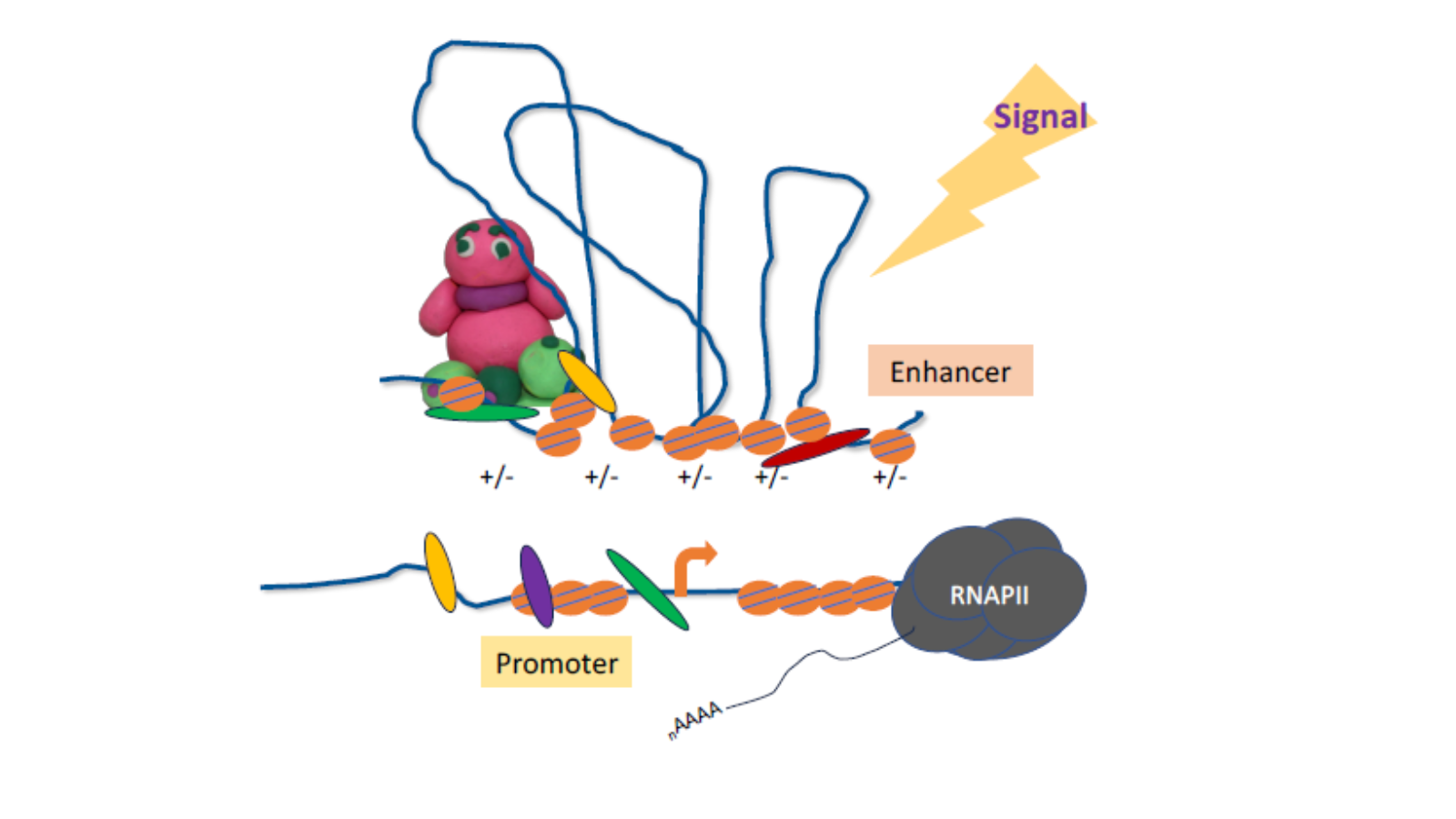
Summary illustration
Key Publications
-
D. Lando, X. Ma, Y. Cao, A. Jartseva, T. J. Stevens, W. Boucher, N. Reynolds, B. Montibus, D. Hall, A Lackner, R. Ragheb, M Leeb, B. D. Hendrich*, E. D. Laue* (2023) bioRxiv Enhancer-promoter interactions are reconfigured through the formation of long-range multiway chromatin hubs as mouse ES cells exit pluripotency https://dx.doi.org/10.1101/2023.02.08.527615 *Co-corresponding
-
Montibus, B., Ragheb, R., Diamanti, E., Dunn, S.-J., Reynolds, N. and Hendrich, B. (2024) The Nucleosome Remodelling and Deacetylation complex coordinates the transcriptional response to lineage commitment in pluripotent cells. Biology Open 13, bio060101 https://doi.org/10.1242/bio.060101
-
Basu, S., Shukron, O., … Klenerman, D*., Hendrich, B.D*., Holcman, D*., Laue, E. D*. (2023) Live-cell three-dimensional single-molecule tracking reveals modulation of enhancer dynamics by NuRD. Nature Structural and Molecular Biology https://doi.org/10.1038/s41594-023-01095-4 *Co-corresponding
-
Ragheb, R., Gharbi, S., Cramard, J., Ogundele, O., Kloet, S. L., Burgold, T., Vermeulen, M., Reynolds, N. and Hendrich, B. (2020). Differential regulation of lineage commitment in human and mouse primed pluripotent stem cells by the nucleosome remodelling and deacetylation complex. Stem Cell Research, 46:101867. PMCID: PMC7347010
- Burgold, T., Barber, M., Kloet, S., Cramard, J., Gharbi, S., Floyd, R., Kinoshita, M., Ralser, M., Vermeulen, M., Reynolds, N., Dietmann, S. and Hendrich, B. (2019). The Nucleosome Remodelling and Deacetylation complex suppresses transcriptional noise during lineage commitment. The EMBO Journal, 38(12):e100788. PMCID: PMC6576150
- Bornelöv S, Reynolds N, Xenophontos M, Gharbi S, Johnstone E, Floyd R, Ralser M, Signolet J, Loos R, Dietmann S, Bertone P, Hendrich B. The Nucleosome Remodeling and Deacetylation Complex Modulates Chromatin Structure at Sites of Active Transcription to Fine-Tune Gene Expression. Mol Cell. 2018 Jul 5;71(1):56-72.e4. PMCID: PMC6039721
- Stevens, T. J., Lando, D., Basu, S., Atkinson, L. P., Cao, Y., Lee, S. F., Leeb, M., Wohlfahrt, K.J., Boucher, W., O’Shaughnessy-Kirwan, A., Cramard, J., Faure, A.J., Ralser, M., Blanco, E., Morey, L., Sansó, M., Palayret, M.G.S., Lehner, B., Di Croce, L., Wutz, A., Hendrich, B., and Laue, E.D. (2017). 3D structures of individual mammalian genomes studied by single-cell Hi-C. Nature, 544(7648) 59-64. doi: 10.1038/nature21429
- Miller, A., Ralser, M., Kloet, S. L., Loos, R., Nishinakamura, R., Bertone, P., Vermeulen, M., and Hendrich, B. (2016). Sall4 controls differentiation of pluripotent cells independently of the Nucleosome Remodelling and Deacetylation (NuRD) complex. Development, 143(17): 3074-84. doi: 10.1242/dev.139113 PMCID:PMC5047675
- O'Shaughnessy-Kirwan A, Signolet J, Costello I, Gharbi S, Hendrich B. (2015). Constraint of gene expression by the chromatin remodelling protein CHD4 facilitates lineage specification. Development, 142(15), 2586–2597. PMCID:PMC4529036
- Reynolds N, Latos P, Hynes-Allen A, Loos R, Leaford D, O’Shaughnessy A, Mosaku O, Signolet J, Brennecke P, Kalkan T, Costello I, Humphreys P, Mansfield W, Nakagawa K, Strouboulis J, Behrens A, Bertone P, Hendrich B. (2012). NuRD suppresses pluripotency gene expression to promote transcriptional heterogeneity and lineage commitment. Cell Stem Cell 10(5): 583-594. PMCID:PMC3402183
- Reynolds N, Salmon-Divon M, Dvinge H, Hynes-Allen A, Balasooriya G, Leaford D, Behrens A, Bertone P, Hendrich B. (2012) NuRD-mediated deacetylation of H3K27 facilitates recruitment of Polycomb Repressive Complex 2 to direct gene repression The EMBO Journal, 31(3), 593–605. PMCID:PMC3273378