Professor Namshik Han

Professor Namshik Han
AI Drug Discovery
Departmental affiliation:
- Milner Therapeutics Institute
- Cambridge Centre for AI in Medicine
- Yonsei University
Email: nh417@cam.ac.uk
Biography
Professor Namshik Han is an internationally recognised scientist at the interface of quantum computing, artificial intelligence, and biomedicine. He leads the AI and Computational Biology group at the Milner Therapeutics Institute, University of Cambridge, where his research focuses on developing quantum algorithms and explainable AI for analysing large-scale multi-modal biomedical data.
At Cambridge, he also serves as Faculty at the Cambridge Centre for AI in Medicine and as an Affiliated Principal Investigator at the Stem Cell Institute, where his work supports translational applications of quantum-AI approaches in stem cell and regenerative medicine. In parallel, he is a Professor in the Department of Quantum Information at Yonsei University, strengthening UK–Korea quantum-AI collaborations.
His lab’s projects span target identification, drug repurposing, patient stratification, and prediction of efficacy and safety, applied across therapeutic areas including cancer, respiratory, CNS, inflammatory, metabolic, and infectious diseases.
Beyond academia, Professor Han is committed to technology translation and entrepreneurship. He was a founding scientist of Storm Therapeutics and co-founded two AI drug discovery startups: CardiaTec Biosciences (AI + human heart tissue multi-omics for cardiovascular disease) and KURE.ai Therapeutics (NK cell-based immuno-oncology).
Research Summary
Professor Han’s group integrates quantum algorithms and AI-driven approaches with multi-modal biomedical data to accelerate therapeutic discovery. The lab develops and applies advanced machine learning, statistical modelling, and network-based methods alongside quantum walk and quantum embedding algorithms to uncover disease pathways, therapeutic targets, and biomarkers.
Research projects span a broad range of disease areas, including cancer, respiratory, central nervous system, inflammatory, metabolic, and infectious diseases. Core activities include:
Therapeutic Target Identification – leveraging multi-omics and network-based quantum-AI to prioritise novel, tractable targets.
Drug Repositioning & Efficacy Prediction – integrating large-scale public datasets with partner-generated experimental data to identify new applications for existing compounds and predict safety/efficacy profiles.
Patient Stratification – using explainable AI and quantum-inspired models to classify patient subgroups and improve personalised treatment strategies.
Mechanistic Discovery – applying causal reasoning and interpretable algorithms to connect computational predictions with molecular mechanisms, enabling validation from in silico to in vitro systems.
The group operates as a single integrated lab across the University of Cambridge and Yonsei University, united under one mission: to pioneer Quantum-AI for next-generation drug discovery. This cross-continental model enables seamless collaboration, resource sharing, and the pursuit of ambitious goals that neither institution could achieve alone.
The lab also leads international efforts to establish Quantum-AI platforms that can transform therapeutic development and global health.
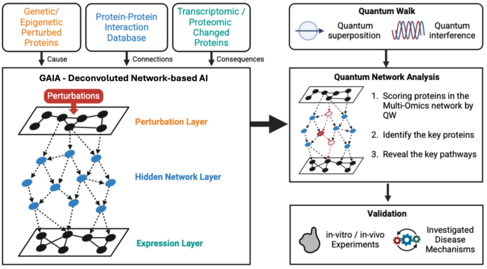
Figure. Quantum-AI network analysis framework
Schematic overview of Professor Han’s research integrating multi-omics data, network-based AI, and quantum walk algorithms for drug discovery. Perturbations at the genetic/epigenetic and transcriptomic/proteomic level are mapped through a deconvoluted network-based AI (GAIA), and scored via quantum network analysis. This approach identifies key proteins and pathways, which are subsequently validated through in vitro and in vivo experiments to investigate disease mechanisms.
CSCI Collaborators
Prof Mekayla Storer
