Professor Ben Simons
Tracing stem cell fate in development, maintenance and disease
Email: bds10@cam.ac.uk
Laboratory: Gurdon Institute
Departmental Affiliation: Applied Mathematics and Theoretical Physics
Biography
Ben has a background in theoretical condensed matter physics. Having obtained his PhD at the Cavendish Laboratory in Cambridge researching high temperature superconductivity, he undertook post-doctoral research in quantum mesoscopic physics at MIT and NEC Research Inc. in Princeton. In 1994 he transferred to a Royal Society Research Fellowship and was appointed to a Lectureship at Imperial College before moving to the Cavendish Laboratory in 1995. In 2002, he was promoted to a Chair in Theoretical Condensed Matter Physics and, in 2011, he was elected to the Herchel Smith Chair in Physics. In 2018 he took up the Royal Society EP Abraham Professorship at the Department of Applied Mathematics and Theoretical Physics and is now also a senior Group Leader at the Wellcome Trust-CRUK Gurdon Institute. His research is supported by grant income from EPSRC, MRC and the Wellcome Trust with whom he holds an Investigator Award.
Funding
EPSRC, MRC, Wellcome Trust
External links
Theory of Condensed Matter page
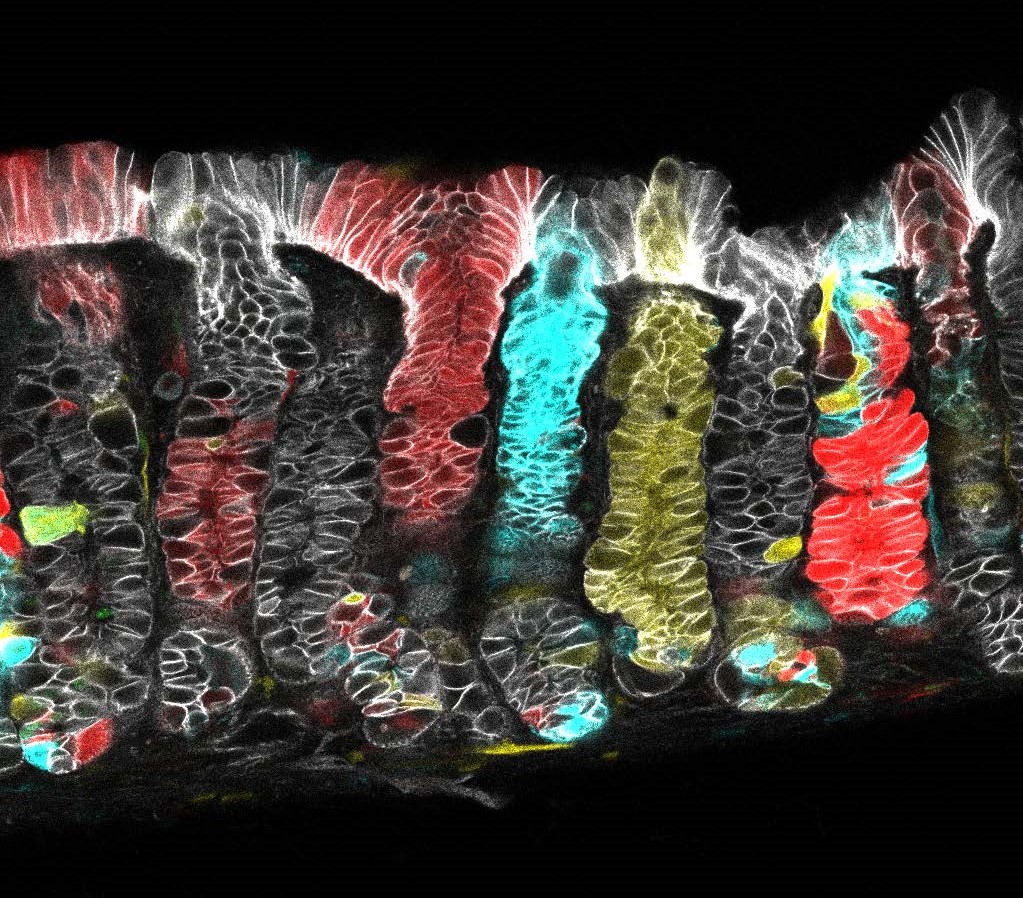
Cell lineage tracing in the stomach corpus (Juergen Fink and Seungmin Han)
Research
Research in the Simons Group combines experimental lineage tracing strategies and single-cell methods with concepts from non-equilibrium statistical physics and mathematics to address the fate behaviour of stem and progenitor cells in the development, maintenance and regeneration of tissues, and factors leading to their dysregulation in diseased states.
In particular, they have resolved strategies of stem cell self-renewal in the maintenance of epithelial tissues, including mammalian brain, epidermis, intestine, lung and testis. Their studies have emphasized the role of stochastic renewal programmes in the regulation of stem cell fate, questioning the nature of stem cell identity and function. They have extended these approaches to study the development and patterning of adult tissues, including the eye, heart, mammary epithelium, pancreas and skin epidermis. At the same, they are collaborating with partner labs to address the cellular basis of tumour initiation. In particular, they are interested in understanding how mechanisms of natural injury-induced cellular reprogramming become subverted during the early stages of tumour development, with ongoing studies in tissues of the gastrointestinal tract, lung, and the brain. Going forward, the group is placing increasing emphasis on the study of developmental programmes in human epithelia, making use of advances in stem cell-derived 3D organ culture models.
Simons Group photo
Plain English
In adults, many tissues such as epidermis, blood, and gut undergo routine and constant turnover. The maintenance and repair of such tissues relies upon stem cells. As with embryonic stem cells, tissue stem cells are defined by their capacity to self-renew and to differentiate into the more specialised cell types. However, in contrast to embryonic stem cells, tissue stem cells must achieve a perfect balance between proliferation and differentiation. Resolving the mechanisms of balance represent one of the defining questions of stem cell biology. To address this question, most studies focus on the identification of molecular regulatory factors. However, such factors are rare and often unspecific. By exploiting methods of population dynamics and statistical physics, we have shown that, in homeostasis, stem cells must follow simple and restricted patterns of fate, which impart characteristic signatures in the size distribution of surviving clones. We are using this general methodology to explore the pattern of tissue maintenance in normal adult tissues, and to address the mechanisms of dysregulation in disease, cancer and aging.
Key Publications
-
M. K. Yum, S. Han, J. Fink, S.-H. Wu, C. Dabrowska, T. Trendafilova, R. Mustata, L. Chatzeli, R. Azzarelli, I. Pshenichnaya, E. Lee, F. England, J. K. Kim, D. E. Stange, A. Philpott, J.-H. Lee, B.-K. Koo and B. D. Simons. Tracing oncogene-driven remodeling of the intestinal stem cell niche. Nature 594, 442-447 (2021) | doi: 10.1038/s41586-021-03605-0
-
S. Han, J. Fink, D. J. Jörg, E. Lee, M. K. Yum, L. Chatzeli, S. R. Merker, M. Josserand, T. Trendafilova, A. Andersson- Rolf, C. Dabrowska, H. Kim, R. Naumann, J.-H. Lee, N. Sasaki, R. L. Mort, I. J. Jackson, O. Basak, H. Clevers, D. E. Stange, A. Philpott, J. K. Kim, B. D. Simons and B.-K. Koo. Defining the identity and dynamics of adult gastric isthmus stem cells. Cell Stem Cell 25, 342-356 (2019) | doi:10.1016/j.stem.2019.07.008
-
Y. Kitadate, D. J. Jörg, M. Tokue, A. Maruyama, R. Ichikawa, S. Tsuchiya, E. Segi-Nishida, T. Nakagawa, A. Uchida, C. Kimura-Yoshida, S. Mizuno, F. Sugiyama, T. Azami, M. Ema, C. Noda, S. Kobayashi, I. Matsuo, Y. Kanai, T. Nagasawa, Y. Sugimoto, S. Takahashi, B. D. Simons and S. Yoshida. Competition for mitogens regulates spermatogenic stem cell homeostasis in an open niche. Cell Stem Cell 24, 79-92 (2019) | doi:10.1016/j.stem.2018.11.013
-
Kitadate Y, Jörg DJ, Tokue M, Maruyama A, Ichikawa R, Tsuchiya S, Segi-Nishida E, Nakagawa T, Uchida A, Kimura-Yoshida C, Mizuno S, Sugiyama F, Azami T, Ema M, Noda C, Kobayashi S, Matsuo I, Kanai Y, Nagasawa T, Sugimoto Y, Takahashi S, Simons BD, Yoshida S. Competition for mitogens regulates spermatogenic stem cell homeostasis in an open niche. Cell Stem Cell (2019) 24, 79-92 PMCID: PMC6327111
- Hannezo E, Scheele CLGJ, Moad M, Drogo N, Heer R, Sampogna RV, van Rheenen J, Simons BD. A unifying theory of branching morphogenesis. Cell (2017) 171, 242-255 PMCID: PMC5610190
- Lan X, Jörg DJ, Cavalli FMG, Richards LM, Nguyen LV, Vanner RJ, Guilhamon P, Lee L, Kushida MM, Pellacani D, Park NI, Coutinho FJ, Whetstone H, Selvadurai HJ, Che C, Luu B, Carles A, Moksa M, Rastegar N, Head R, Dolma S, Prinos P, Cusimano MD, Das S, Bernstein M, Arrowsmith CH, Mungall AJ, Moore RA, Ma Y, Gallo M, Lupien M, Pugh TJ, Taylor MD, Hirst M, Eaves CJ, Simons BD, Dirks PB. Fate mapping of human glioblastoma reveals an invariant stem cell hierarchy. Nature (2017) 549, 227-232 PMCID: PMC5608080
- Sánchez-Danés A, Hannezo E, Larsimont JC, Liagre M, Youssef KK, Simons BD, Blanpain C. Defining the clonal dynamics leading to mouse skin tumour initiation. Nature (2016) 536, 298-303 PMCID: PMC5068560
- Simons BD, Clevers H. Strategies for homeostatic stem cell self-renewal in adult tissues. Cell (2011). 145, 851-862. PMID:21663791